
NMR Restraints Grid
 |
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | cing | stage | position | type |
|
|
6198 |
1iml |
cing | 1-original | 1 | comment |
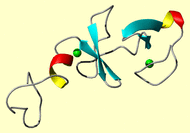
Experimental Restraints for the NMR Structures of the
LIM Only protein Cysteine Rich Intestinal Protein, CRIP
The solution NMR 3D structure of the Cysteine Rich Intestinal Protein,
CRIP, is based on 500 distance restraints derived
from the NOE data and 17 metal restraints. Hydrogen bond distance restraints
were not used for generation and refinement of the last 12 MODELS (MODEL 37-48).
A complete list of experimental restraints follows:
List of residues with corresponding numbering in the sequence
of the 48 MODELS:
1 PKCPKCDKEV YFAERVTSLG
21 KDWHRPCLKC EKCGKTLTSG
41 GHAEHEGKPY CNHPCYSAMP
61 GPKGFGRGGA ESHTFK
Pro-1 Lys-2 Cys-3 Pro-4 Lys-5 Cys-6 Asp-7 Lys-8 Glu-9 Val-10
Tyr-11 Phe-12 Ala-13 Glu-14 Arg-15 Val-16 Thr-17 Ser-18 Leu-19 Gly-20
Lys-21 Asp-22 Trp-23 His-24 Arg-25 Pro-26 Cys-27 Leu-28 Lys-29 Cys-30
Glu-31 Lys-32 Cys-33 Gly-34 Lys-35 Thr-36 Leu-37 Thr-38 Ser-39 Gly-40
Gly-41 His-42 Ala-43 Glu-44 His-45 Glu-46 Gly-47 Lys-48 Pro-49 Tyr-50
Cys-51 Asn-52 His-53 Pro-54 Cys-55 Tyr-56 Ser-57 Ala-58 Met-59 Phe-60
Gly-61 Pro-62 Lys-63 Gly-64 Phe-65 Gly-66 Arg-67 Gly-68 Gly-69 Ala-70
Glu-71 Ser-72 His-73 Thr-74 Phe-75 Lys-76
NOE Interproton distance restraints. Distances are in Angstroms (A)
Strong 1.8 - 2.7 A
Medium 1.8 - 3.3 A
Weak 1.8 - 5.0 A
Restraints to dummy atoms:
+ 0.5 for -CH3
+ 2.13 for cg-cd-hd of Tyr and Phe
+ 2.13 for cz-ce-he of Tyr
+ 2.14 for cz-ce-he of Phe
Metal restraints used to enforce Zn-S and Zn-N bond distances,
and to ensure proper hybridization of the His-Nd and Cys-Sg atoms
(distances in A)
LB= Lower Bound
UB= Upper Bound
ATOM 1 ATOM 2 distance
Contact the webmaster for help, if required. Tuesday, June 18, 2024 1:35:52 PM GMT (wattos1)