
NMR Restraints Grid
 |
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | bmrb_id | cing | stage | program | type |
|
|
37000 |
1osl |
5791 | cing | 2-parsed | STAR | comment |
data_1osl_MR_file_constraints
save_Conversion_project
_Study_list.Sf_category study_list
_Study_list.Entry_ID parsed_1osl
_Study_list.ID 1
loop_
_Study.ID
_Study.Name
_Study.Type
_Study.Details
_Study.Entry_ID
_Study.Study_list_ID
1 "Conversion project" NMR . parsed_1osl 1
stop_
save_
save_entry_information
_Entry.Sf_category entry_information
_Entry.ID parsed_1osl
_Entry.Title "Original constraint list(s)"
_Entry.Version_type original
_Entry.Submission_date .
_Entry.Accession_date .
_Entry.Last_release_date .
_Entry.Original_release_date .
_Entry.Origination .
_Entry.NMR_STAR_version 3.1
_Entry.Original_NMR_STAR_version .
_Entry.Experimental_method NMR
_Entry.Experimental_method_subtype .
loop_
_Related_entries.Database_name
_Related_entries.Database_accession_code
_Related_entries.Relationship
_Related_entries.Entry_ID
PDB 1osl "Master copy" parsed_1osl
stop_
save_
save_global_Org_file_characteristics
_Constraint_stat_list.Sf_category constraint_statistics
_Constraint_stat_list.Entry_ID parsed_1osl
_Constraint_stat_list.ID 1
loop_
_Constraint_file.ID
_Constraint_file.Constraint_filename
_Constraint_file.Software_ID
_Constraint_file.Software_label
_Constraint_file.Software_name
_Constraint_file.Block_ID
_Constraint_file.Constraint_type
_Constraint_file.Constraint_subtype
_Constraint_file.Constraint_subsubtype
_Constraint_file.Constraint_number
_Constraint_file.Entry_ID
_Constraint_file.Constraint_stat_list_ID
1 1osl.mr . . "MR format" 1 comment "Not applicable" "Not applicable" 0 parsed_1osl 1
1 1osl.mr . . XPLOR/CNS 2 distance NOE simple 0 parsed_1osl 1
1 1osl.mr . . XPLOR/CNS 3 distance "hydrogen bond" simple 0 parsed_1osl 1
1 1osl.mr . . XPLOR/CNS 4 distance NOE simple 0 parsed_1osl 1
1 1osl.mr . . XPLOR/CNS 5 "dihedral angle" "Not applicable" "Not applicable" 0 parsed_1osl 1
1 1osl.mr . . "MR format" 6 "nomenclature mapping" "Not applicable" "Not applicable" 0 parsed_1osl 1
stop_
save_
save_MR_file_comment_1
_Org_constr_file_comment.Sf_category org_constr_file_comment
_Org_constr_file_comment.Entry_ID parsed_1osl
_Org_constr_file_comment.ID 1
_Org_constr_file_comment.Constraint_file_ID 1
_Org_constr_file_comment.Block_ID 1
_Org_constr_file_comment.Details "Generated by Wattos"
_Org_constr_file_comment.Comment
;
*HEADER TRANSCRIPTION/DNA 20-MAR-03 1OSL
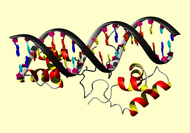
*TITLE SOLUTION STRUCTURE OF A DIMERIC LACTOSE DNA-BINDING DOMAIN
*TITLE 2 COMPLEXED TO A NONSPECIFIC DNA SEQUENCE
*COMPND MOL_ID: 1;
*COMPND 2 MOLECULE: LACTOSE OPERON REPRESSOR;
*COMPND 3 CHAIN: A, B;
*COMPND 4 FRAGMENT: N-TERMINAL DNA-BINDING DOMAIN, RESIDUES 1-62;
*COMPND 5 ENGINEERED: YES;
*COMPND 6 MUTATION: YES;
*COMPND 7 MOL_ID: 2;
*COMPND 8 MOLECULE: 5'-
*COMPND 9 D(*CP*GP*AP*TP*AP*AP*GP*AP*TP*AP*TP*CP*TP*TP*AP*TP*CP*G)-
*COMPND 10 3';
*COMPND 11 CHAIN: C, D;
*COMPND 12 ENGINEERED: YES
*SOURCE MOL_ID: 1;
*SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
*SOURCE 3 ORGANISM_COMMON: BACTERIA;
*SOURCE 4 GENE: LACI;
*SOURCE 5 EXPRESSION_SYSTEM: ESCHERICHIA COLI;
*SOURCE 6 EXPRESSION_SYSTEM_COMMON: BACTERIA;
*SOURCE 7 EXPRESSION_SYSTEM_STRAIN: DH9;
*SOURCE 8 EXPRESSION_SYSTEM_VECTOR_TYPE: PLASMID;
*SOURCE 9 EXPRESSION_SYSTEM_PLASMID: PET-HP62-V52C;
*SOURCE 10 MOL_ID: 2;
*SOURCE 11 SYNTHETIC: YES
*KEYWDS PROTEIN-DNA COMPLEX, LAC REPRESSOR, NONSPECIFIC INTERACTION
*EXPDTA NMR, 20 STRUCTURES
*AUTHOR C.KALODIMOS, A.BONVIN, R.BOELENS, R.KAPTEIN
*REVDAT 1 04-MAY-04 1OSL 0
;
save_
Contact the webmaster for help, if required. Friday, May 3, 2024 10:17:39 AM GMT (wattos1)