
NMR Restraints Grid
 |
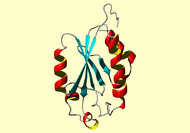
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | bmrb_id | cing | stage | program | type |
|
|
13624 |
1v9w |
10080 | cing | 1-original | MR format | comment |
*HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 03-FEB-04 1V9W *TITLE SOLUTION STRUCTURE OF MOUSE PUTATIVE 42-9-9 PROTEIN *COMPND MOL_ID: 1; *COMPND 2 MOLECULE: PUTATIVE 42-9-9 PROTEIN; *COMPND 3 CHAIN: A; *COMPND 4 ENGINEERED: YES *SOURCE MOL_ID: 1; *SOURCE 2 ORGANISM_SCIENTIFIC: MUS MUSCULUS; *SOURCE 3 ORGANISM_COMMON: HOUSE MOUSE; *SOURCE 4 ORGANISM_TAXID: 10090; *SOURCE 5 GENE: RIKEN CDNA 1110050H17; *SOURCE 6 EXPRESSION_SYSTEM_VECTOR_TYPE: PLASMID; *SOURCE 7 EXPRESSION_SYSTEM_PLASMID: P020128-01; *SOURCE 8 OTHER_DETAILS: CELL-FREE PROTEIN SYNTHESIS *KEYWDS THIOREDOXIN-LIKE FOLD, STRUCTURAL GENOMICS, RIKEN *KEYWDS 2 STRUCTURAL GENOMICS/PROTEOMICS INITIATIVE, RSGI, UNKNOWN *KEYWDS 3 FUNCTION *EXPDTA SOLUTION NMR *NUMMDL 20 *AUTHOR N.TOCHIO, S.KOSHIBA, M.INOUE, T.KIGAWA, S.YOKOYAMA, RIKEN *AUTHOR 2 STRUCTURAL GENOMICS/PROTEOMICS INITIATIVE (RSGI) *REVDAT 1 12-MAY-09 1V9W 0 ************************************************************** During the CYANA calculations automatic implicit swapping of restraints involving diastereotopic substitutents was applied for prochrial groups without stereospecific assignment. Diastereotopic substitents were swapped individually in each conformer to calculate the minimal target function and restraint violations. The optimal swapping for a given prochiral group may differ among the 20 conformers that represent the solution structure. The swapping is therefore performed implicitly in the program and is not reflected in the distance restraint file deposited in the PDB. **************************************************************
Contact the webmaster for help, if required. Friday, June 14, 2024 10:31:54 PM GMT (wattos1)