
NMR Restraints Grid
 |
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | bmrb_id | cing | stage | position | program | type |
|
|
11601 |
1rqu |
4429 | cing | 1-original | 9 | DYANA/DIANA | coupling constant |
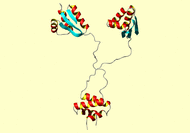
#1H-15N RDC orientation constraints for structure calculation of L7 CTD in virus alignment medium 53 GLU- HN 3.770 0.089 54 PHE HN 2.400 0.121 55 ASP- HN 2.820 0.122 56 VAL HN -1.280 0.081 57 ILE HN 0.760 0.112 58 LEU HN -4.070 0.067 59 LYS+ HN -3.580 0.064 60 ALA HN -3.250 0.085 61 ALA HN -3.500 0.071 62 GLY HN -0.550 0.076 63 ALA HN -0.600 0.130 65 LYS+ HN -2.280 0.086 66 VAL HN -3.310 0.094 67 ALA HN -3.840 0.086 68 VAL HN -3.250 0.058 69 ILE HN -1.920 0.133 70 LYS+ HN -3.880 0.081 71 ALA HN -4.050 0.064 72 VAL HN -2.350 0.089 73 ARG+ HN -2.190 0.089 74 GLY HN -3.830 0.064 75 ALA HN -3.750 0.094 76 THR HN 0.690 0.086 77 GLY HN -3.440 0.064 78 LEU HN 1.790 0.112 79 GLY HN -2.100 0.112 80 LEU HN 4.060 0.103 81 LYS+ HN 0.700 0.098 82 GLU- HN -0.140 0.092 83 ALA HN 4.610 0.108 84 LYS+ HN 3.370 0.086 85 ASP- HN 0.380 0.086 86 LEU HN 1.480 0.064 87 VAL HN 4.500 0.076 88 GLU- HN 1.140 0.078 89 SER HN -2.300 0.081 90 ALA HN 5.050 0.058 92 ALA HN -3.350 0.092 93 ALA HN -3.110 0.114 94 LEU HN 1.560 0.086 95 LYS+ HN 2.020 0.100 96 GLU- HN 0.960 0.058 97 GLY HN -0.870 0.104 98 VAL HN -2.840 0.071 99 SER HN 4.010 0.136 100 LYS+ HN 2.200 0.122 101 ASP- HN -1.540 0.112 102 ASP- HN 0.850 0.148 103 ALA HN 3.170 0.148 104 GLU- HN -0.370 0.085 105 ALA HN -0.970 0.155 106 LEU HN 2.120 0.086 107 LYS+ HN 1.850 0.112 108 LYS+ HN -1.530 0.076 109 ALA HN -1.740 0.071 110 LEU HN 2.760 0.130 111 GLU- HN -0.490 0.114 112 GLU- HN -2.850 0.058 113 ALA HN 0.020 0.078 114 GLY HN 0.220 0.064 115 ALA HN -2.740 0.094 116 GLU- HN -1.660 0.078 117 VAL HN -2.950 0.086 118 GLU- HN -0.590 0.100 119 VAL HN -1.290 0.058 120 LYS+ HN 2.570 0.092
Contact the webmaster for help, if required. Sunday, May 19, 2024 2:34:13 PM GMT (wattos1)