
NMR Restraints Grid
 |
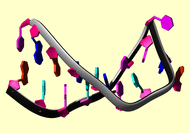
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | cing | stage | position | program | type | subtype | subsubtype |
|
|
468012 |
2kpd |
cing | 1-original | 2 | XPLOR/CNS | distance | hydrogen bond | simple |
!A17-U 1
assign (resid 17 and name N1)
(resid 1 and name N3)
2.82 0.6 0.6
assign (resid 17 and name N6)
(resid 1 and name O4)
2.95 0.6 0.6
!A 3-U15
assign (resid 3 and name N1)
(resid 15 and name N3)
2.82 0.6 0.6
assign (resid 3 and name N6)
(resid 15 and name O4)
2.95 0.6 0.6
!G 2-C16
assign (resid 2 and name N1)
(resid 16 and name N3)
2.95 0.6 0.6
assign (resid 2 and name O6)
(resid 16 and name N4)
2.91 0.6 0.6
assign (resid 2 and name N2)
(resid 16 and name O2)
2.86 0.6 0.6
!G 4-C14
assign (resid 4 and name N1)
(resid 14 and name N3)
2.95 0.6 0.6
assign (resid 4 and name O6)
(resid 14 and name N4)
2.91 0.6 0.6
assign (resid 4 and name N2)
(resid 14 and name O2)
2.86 0.6 0.6
!G13-C5
assign (resid 13 and name N1)
(resid 5 and name N3)
2.95 0.6 0.6
assign (resid 13 and name O6)
(resid 5 and name N4)
2.91 0.6 0.6
assign (resid 13 and name N2)
(resid 5 and name O2)
2.86 0.6 0.6
!C7-U11 added based on 13C chemical shifts
assign (resid 7 and name N3)
(resid 11 and name O4)
2.82 1.0 1.0
assign (resid 7 and name O2)
(resid 11 and name N3)
2.95 1.0 1.0
end
set message=on echo=on end
Contact the webmaster for help, if required. Thursday, July 4, 2024 12:52:28 AM GMT (wattos1)