
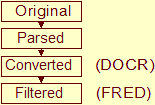
NMR Restraints Grid
 |
NMR Restraints Grid |

|
Result table
| image | mrblock_id | pdb_id | bmrb_id | cing | stage | position | program | type | subtype | subsubtype | item_count | other_prop |
|
|
2174 |
1bhu |
4217 | cing | 1-original | 0 | MR format | comment |
|
|
|
|
|
|
2175 |
1bhu |
4217 | cing | 1-original | 1 | XPLOR/CNS | distance | NOE | simple |
|
|
|
|
2176 |
1bhu |
4217 | cing | 1-original | 2 | XPLOR/CNS | distance | hydrogen bond | simple |
|
|
|
|
2177 |
1bhu |
4217 | cing | 1-original | 3 | XPLOR/CNS | distance | NOE | simple |
|
|
|
|
2178 |
1bhu |
4217 | cing | 1-original | 4 | XPLOR/CNS | distance | disulfide bond | simple |
|
|
|
|
2179 |
1bhu |
4217 | cing | 1-original | 5 | XPLOR/CNS | dihedral angle |
|
|
|
|
|
|
2180 |
1bhu |
4217 | cing | 1-original | 6 | MR format | nomenclature mapping |
|
|
|
|
|
|
29872 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | comment |
|
|
0 |
|
|
|
29873 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | distance | NOE | simple | 1443 |
|
|
|
29874 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | distance | hydrogen bond | simple | 64 |
|
|
|
29875 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | distance | NOE | simple | 23 |
|
|
|
29876 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | distance | disulfide bond | simple | 6 |
|
|
|
29877 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | dihedral angle |
|
|
52 |
|
|
|
29878 |
1bhu |
4217 | cing | 2-parsed | 0 | STAR | entry | full |
|
1588 |
|
|
|
369887 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | STAR | entry | full |
|
1588 |
|
|
|
369888 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XML | entry | full |
|
|
|
|
|
369889 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | sequence |
|
|
|
|
|
|
369890 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | coordinate | ensemble |
|
|
|
|
|
369891 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | distance | hydrogen bond | ambi |
|
|
|
|
369892 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | distance | general distance | ambi |
|
|
|
|
369893 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | distance | general distance | ambi |
|
|
|
|
369894 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | distance | general distance | ambi |
|
|
|
|
369895 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | dihedral angle |
|
|
|
|
|
|
369896 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | sequence |
|
|
|
|
|
|
369897 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | hydrogen bond | ambi |
|
|
|
|
369898 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
|
|
|
369899 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
|
|
|
369900 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
|
|
|
369901 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
LOWER_ONLY=true |
|
|
369902 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
LOWER_ONLY=true |
|
|
369903 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
LOWER_ONLY=true |
|
|
369904 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | hydrogen bond | ambi |
|
LOWER_ONLY=true |
|
|
369905 |
1bhu |
4217 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | dihedral angle |
|
|
|
|
|
|
369906 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | STAR | entry | full |
|
1536 |
|
|
|
369907 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | Wattos | check | stereo assignment | distance |
|
|
|
|
369908 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | Wattos | check | surplus | distance |
|
|
|
|
369909 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | Wattos | check | violation | distance |
|
|
|
|
369910 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | Wattos | check | violation | dihedral angle |
|
|
|
|
369911 |
1bhu |
4217 | cing | 4-filtered-FRED | 0 | Wattos | check | completeness | distance |
|
|
Contact the webmaster for help, if required. Monday, May 13, 2024 9:03:15 AM GMT (wattos1)